The problem
Wearable sensors pick up everything. Your heartbeat, yes, but also every step you take, every arm swing, every fidget. The signal you actually want gets buried under motion artifacts, baseline drift, and noise. If you don't clean it before analysis, your heart rate estimates are wrong, your HRV features are meaningless, and everything downstream breaks. Especially with kids... they move a lot.
This pipeline is my attempt to fix that.
What I built
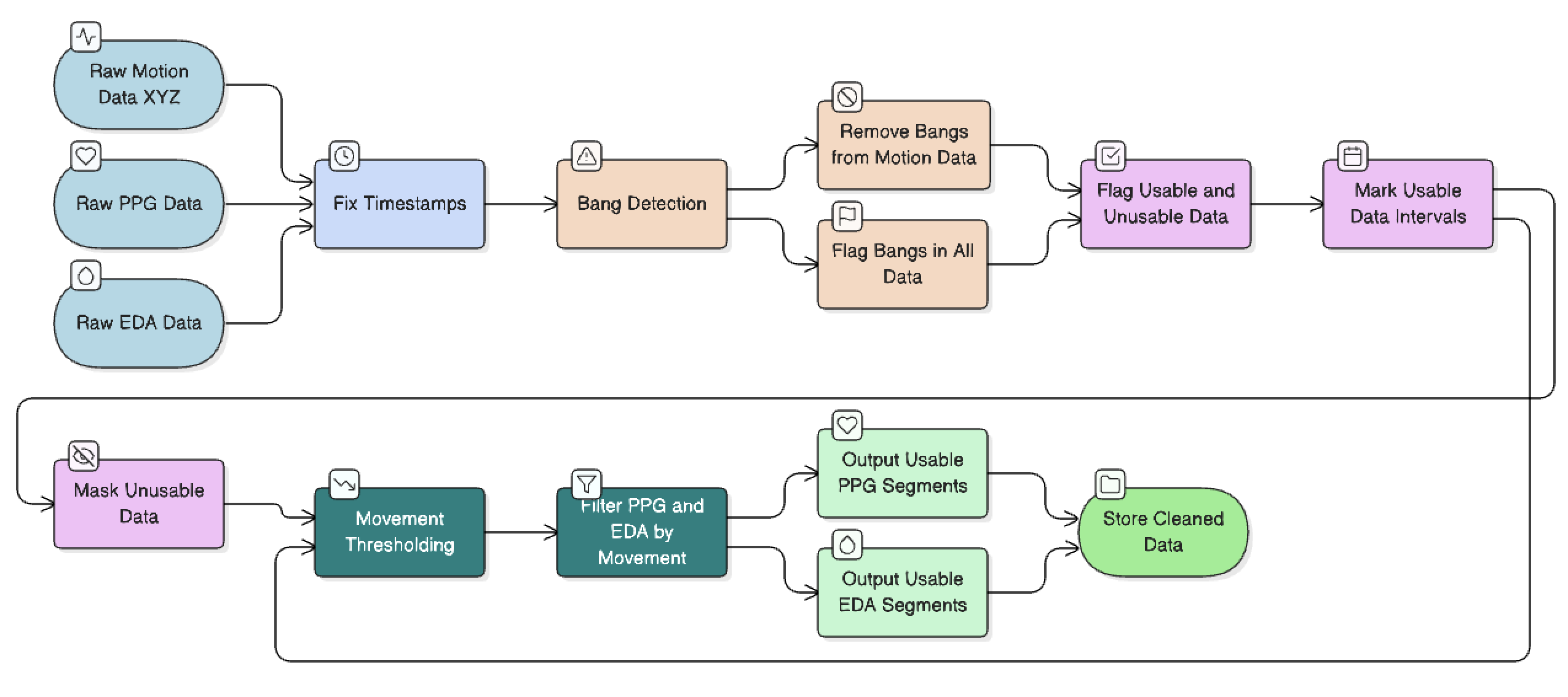
The pipeline ingests raw PPG and EDA data from wearable sensors and runs it through a series of processing stages. The interesting part was motion artifact removal. Hard sudden movements I called "bangs" leave a distinct signature across the accelerometer axes, and once I found it, I could flag and remove those windows automatically before they could contaminate the downstream features.
What I found
Motion artifacts are the biggest source of error in wrist-worn PPG, and they're not random, there's usually a pattern.
Want to know more?
I can't share the full dataset or codebase publicly, but I'm happy to talk through the methods, the design decisions, or the messier parts of real-world biosensor data collection. If any of this is relevant to what you're working on, reach out →